Saad Ahmad
I build AI systems that move from research notebooks to usable products — spanning deep learning, medical image analysis, backend inference APIs, and interactive web interfaces.

Research
Master's Thesis
FAU Erlangen-Nuremberg · Smart Imaging Lab · Supervisor: Prof. Dr. Jana Hutter
Thesis I · 17 September 2025 - 17 March 2026
Automatic lumbosacral vertebra localization in variable field-of-view pelvic MRI
4.21 mm
Mean localization error
S1-S5
Individual sacral detection
1.4s
Per-volume inference
Multi-Center
MRI protocol robustness
Problem Statement
Automatic vertebra localization in pelvic MRI is challenging due to variable field-of-view coverage, anatomical diversity, and partial visibility of the lumbosacral spine. Manual landmark annotation is time-consuming and subject to inter-observer variability, limiting reproducibility in clinical workflows.
Approach
Developed a dual-head 3D U-Net for volumetric vertebra landmark detection in sagittal T2-weighted pelvic MRI. The system jointly predicts vertebral center heatmaps and an anatomical S1 reference point, enabling robust S1-anchored labeling across incomplete or variable spine coverage.
Results
Achieved a mean vertebral localization error of 4.21 mm across multi-center pelvic MRI datasets, with robust detection of lumbar and sacral vertebrae (L1-L5, S1-S5).
Model output · live inference
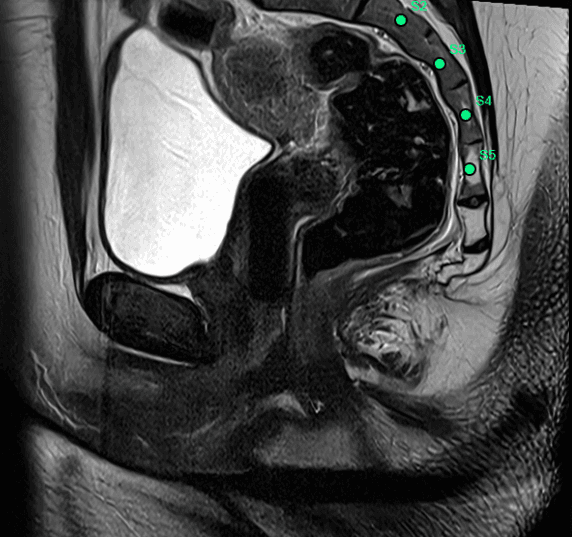
Thesis II · 17 September 2025 - 17 March 2026
Automatic detection of uterine anatomical landmarks in pelvic MRI
4.52 mm
Overall mean error
192 landmarks evaluated
2.90 mm
Best landmark precision
Cavity Cervix
3.66 mm
Median localization error
All 6 landmarks
32
Multi-protocol test cases
3 acquisition protocols
Problem Statement
Uterine biometry — measuring fundal thickness, body length, cervical length, and AP diameter — is essential for managing endometrial cancer, fibroids, and endometriosis, yet is performed manually with high inter-observer variability. No prior automated 3D approach existed for multi-landmark localization.
Approach
Built the preprocessing pipeline used across the entire project, handling raw MRI data from three different scanner protocols. Using an nnU-Net v2 uterine segmentation as a region of interest, trained a multi-decoder 3D U-Net with a shared encoder and independent decoder branches for each landmark group, predicting six uterine landmarks simultaneously. Introduced SharpHeatmapLoss — a custom loss combining MSE with a cubic penalty term that forces sharply peaked heatmaps — and ROI-guided inference using uterine segmentation masks to eliminate background noise.
Results
Achieved 4.52 mm overall mean error across 192 landmarks and 32 test cases from 3 acquisition protocols. Cavity Cervix and Cavity Fundus reached sub-3 mm precision. Landmark detection contributed directly to a real-time reporting pipeline achieving end-to-end results in under 60 seconds, submitted to IEEE Transactions on Medical Imaging.
Model output · real predictions
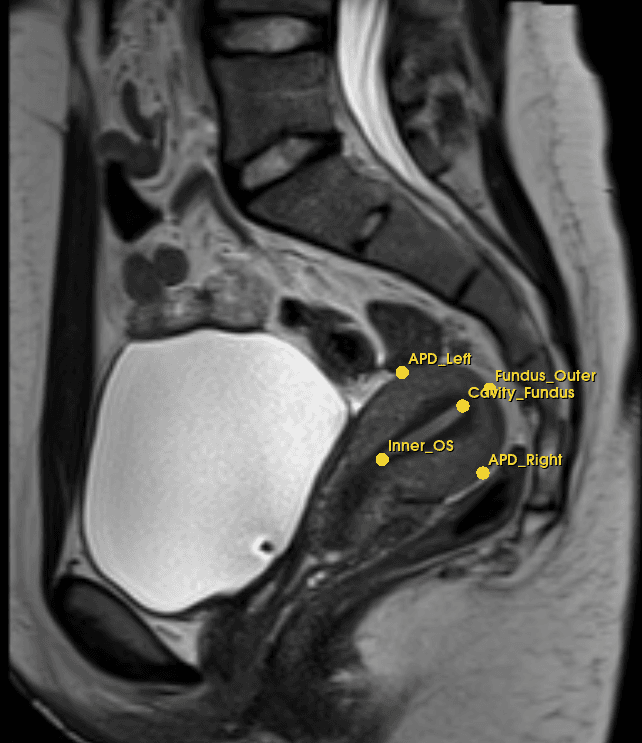
Career
Work Experience
AI / ML Engineer
CurrentYour Company
Jan 2024 – Present
Remote
- Developed and deployed a 3D U-Net pipeline for vertebral landmark detection, serving live predictions via FastAPI.
- Reduced inference latency by 40% through model quantisation and async batching.
- Built end-to-end CI/CD pipeline with GitHub Actions and Docker for zero-downtime model updates.
Full-Stack Developer
Previous Company
Jun 2022 – Dec 2023
On-site
- Architected and shipped React/Next.js web applications serving 10k+ monthly active users.
- Designed RESTful APIs with FastAPI and PostgreSQL; implemented JWT auth and role-based access.
- Mentored junior developers and led weekly code-review sessions.
Research Assistant — Medical Imaging
Research Lab / University
Sep 2021 – May 2022
University
- Annotated and pre-processed 400+ sacral MRI volumes for deep learning dataset construction.
- Implemented baseline atlas-based landmark detection methods for benchmark comparison.
Selected work